Coronavirus Mutation Concern Verified by Largest COVID-19 Viral Sequence Analysis in U.S.
By
By
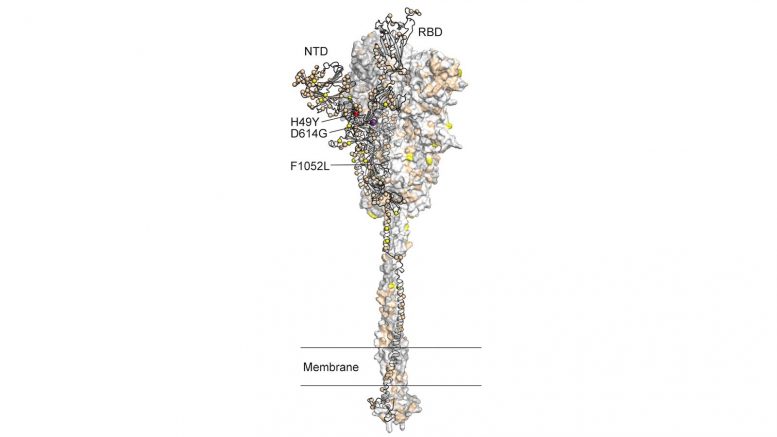 Argonne computational resources supported the largest comprehensive analysis of COVID-19 genome sequences in the U.S. and helped corroborate growing evidence of a protein mutation.
Argonne computational resources supported the largest comprehensive analysis of COVID-19 genome sequences in the U.S. and helped corroborate growing evidence of a protein mutation.Before COVID-19 first entered the United States in March, Houston Methodist Hospital had already begun preparations to test for and sequence the virus on a large scale, given the news coming out of Wuhan, China.
Between March 5, when the first case turned up in metropolitan Houston, and July 7, physician/researchers at Houston Methodist sequenced the genome of over 5,085 strains of the virus.
These accounted for nearly 10 percent of the COVID-19 cases that came through the 2,400-bed Houston Methodist health system, during two distinct waves that occurred in that time frame.
“99 percent isn’t 100 percent. If there is a mutation that accounts for just one percent of the population, and you suppress or eradicate the majority, you can drive up some trait of that one percent, whether it’s virulence or transmissibility, and then it’s a different ballgame.” — James Davis, Argonne staff scientist
“99 percent isn’t 100 percent. If there is a mutation that accounts for just one percent of the population, and you suppress or eradicate the majority, you can drive up some trait of that one percent, whether it’s virulence or transmissibility, and then it’s a different ballgame.” — James Davis, Argonne staff scientist
Collaborators from the University of Texas at Austin, Weill Cornell Medical College, the University of Chicago and the U.S. Department of Energy’s (DOE) Argonne National Laboratory worked together to analyze the data and try to correlate patient outcomes with viral traits.
“This is the largest viral sequence analysis in the U.S. right now and it’s one of the most comprehensive, continual snapshots of sequences that dates to the beginning of the outbreak,” said James Davis, a staff scientist in Argonne’s Data Science and Learning division. “It also provides a much clearer picture of how the strains are evolving.”
During the course of the research, the group helped solidify mounting observations and concerns internationally that a mutation in the virus’s spike protein had become dominant, driving COVID-19’s transmissibility rates as witnessed by the second wave that surged through Houston around mid-May.
A paper describing their methods and results was published in the journal mBio on October 30 ,2020.
That mutation in the spike — responsible for infiltrating the human immune system and the current target of vaccine research — was in an amino acid called Gly614 and was the result of one protein, aspartic acid, mutating into another, glycine.
A paper describing their methods and results was published in the journal mBio on October 30 ,2020.
That mutation in the spike — responsible for infiltrating the human immune system and the current target of vaccine research — was in an amino acid called Gly614 and was the result of one protein, aspartic acid, mutating into another, glycine.
During the earliest parts of the pandemic, March through April, Gly614 was just one variant among many others. But during the second wave in May, Davis recalls, all of the cases they were sequencing at Houston Methodist showed that Gly614 had proliferated to the point of becoming the dominant amino acid in the spike protein.
In fact, it was found in over 99 percent of the sequenced variants.
“The SARS-CoV-2 virus is remarkably conserved, so whenever you see changes like this it’s more remarkable because you don’t tend to see that many mutations,” he said. “I’m not sure if it makes the virus more virulent or easier to transmit, but the study does show some data suggesting that patients with the Gly614 mutation have a larger viral load, though they aren’t necessarily sicker.”
Coincident with the Gly614 takeover in the second wave, patients tended to be younger, showed less severe symptoms, were more likely to be Hispanic/Latino, and lived in areas of lower median incomes. Still, the reasons were unclear and they’d hoped that Argonne’s computational resources would open a door on the causes.
A working relationship already in place, Houston Methodist approached Argonne for help with genome sequence analyses of the 5,000-plus COVID strains, as well as phylogenetic analyses, which look at changes in an organism or a specific feature over time.
Through its Bioinformatics Resource Center Project, supported by the National Institute of Allergy and Infectious Diseases, Argonne provides computational and technical resources to collaborators conducting large biological-based data projects. In this case, it managed the sequencing components, and included quality control, genome alignments and the building of phylogenetic trees.
“For a virus, it has a pretty big genome,” noted Davis, “so, the process became computationally expensive.” But with its arsenal of computers, both large and super, Argonne was prepared to handle the influx of data.
Another aspect of that work involved the artificial intelligence technique called machine learning.
While the focus of machine learning among many research institutions, including Argonne, has focused on determining how drugs might interact with COVID-19, Marcus Nguyen hoped to predict whether the sequence of the virus could ultimately predict patient outcome or patient demographics.
Nguyen, a research specialist with a joint appointment at Argonne and the University of Chicago, looked at correlations between the genome sequences and patient information.
Nguyen, a research specialist with a joint appointment at Argonne and the University of Chicago, looked at correlations between the genome sequences and patient information.
The process trained a machine learning algorithm on the genomic sequences from Houston Methodist, as well as past patient outcomes — length of hospital stay, need for mechanical ventilation, mortality — to potentially determine those outcomes.
“Unfortunately, we didn’t get the results we were hoping for,” said Nguyen. “Although there have been a few correlations found in different pieces of patient metadata, I don’t think there is anything in the genome that is indicative of patient outcome, so there has to be something else going on.”
While the group did observe other variations in the spike, research continues on Gly614 to understand its dynamics and determine what role, if any, it might play in treatment therapies.
“It should be helpful in vaccine development because it shows so many different sequences and so many variants that you get a pretty good picture of what to look for in terms of what variants are dominant in the population,” said Davis.
Though the virus has remained somewhat stable, to date, this study — and natural history — have shown that it only takes one mutation to create a powerful impact on life, both large and diminutive.
“99 percent isn’t 100 percent,” Davis pointed out. “If there is a mutation that accounts for just one percent of the population, and you suppress or eradicate the majority, you can drive up some trait of that one percent, whether it’s virulence or transmissibility, and then it’s a different ballgame.”
“99 percent isn’t 100 percent,” Davis pointed out. “If there is a mutation that accounts for just one percent of the population, and you suppress or eradicate the majority, you can drive up some trait of that one percent, whether it’s virulence or transmissibility, and then it’s a different ballgame.”
This research appears in the article “Molecular Architecture of Early Dissemination and Massive Second Wave of the SARS-CoV-2 Virus in a Major Metropolitan Area,” in mBio, October 30, 2020. Maulik Shukla, of both Argonne and the University of Chicago, was among the co-authors.
Read Coronavirus Genetic Mutation May Have Made COVID-19 More Contagious for more on this research.
Read Coronavirus Genetic Mutation May Have Made COVID-19 More Contagious for more on this research.
Reference: “Molecular Architecture of Early Dissemination and Massive Second Wave of the SARS-CoV-2 Virus in a Major Metropolitan Area” by S. Wesley Long, Randall J. Olsen, Paul A. Christensen, David W. Bernard, James J. Davis, Maulik Shukla, Marcus Nguyen, Matthew Ojeda Saavedra, Prasanti Yerramilli, Layne Pruitt, Sishir Subedi, Hung-Che Kuo, Heather Hendrickson, Ghazaleh Eskandari, Hoang A. T. Nguyen, J. Hunter Long, Muthiah Kumaraswami, Jule Goike, Daniel Boutz, Jimmy Gollihar, Jason S. McLellan, Chia-Wei Chou, Kamyab Javanmardi, Ilya J. Finkelstein and James M. Musser, 30 October 2020, mBio.
DOI: 10.1128/mBio.02707-20
Research funding was provided by the National Institutes of Health (NIH) National Institute of Allergy and Infectious Diseases (NIAID) Bacterial and Viral Bioinformatics resource center.




No comments:
Post a Comment